In many real-world inverse problems, a forward model depends on several fundamentally distinct physical quantities or parameters, rather than a single, homogeneous set of variables. For instance, in a mechanical problem, one might simultaneously infer material stiffness, boundary conditions, and external loads. These parameters often have different physical units, vastly distinct numerical scales, and completely different prior beliefs. While mathematically possible, grouping these distinct quantities into a single long vector can obscure their individual semantic properties, lead to poor inference, and make constructing customized, efficient Markov Chain Monte Carlo (MCMC) algorithms exceedingly difficult.
CUQIpy provides an elegant and mathematically rigorous solution to this challenge through its native support for multiple-input models and Gibbs sampling strategy. By allowing users to define separate geometries and prior distributions for each input element/group, developers can build a multiple-input forward Model that respects the individual nature of each input element/group. When paired with the HybridGibbs sampler, this allows users to apply tailored, element/group-wise sampling strategies, such as using a Metropolis-Hastings (MH) step for lower-dimensional groups where gradient information is difficult to obtain, while employing the Metropolis-Adjusted Langevin Algorithm (MALA) for high-dimensional ones where gradient is available.
To intuitively demonstrate this capability, we will explore a very simplified “bathtub” mixing problem. Imagine a bathtub filled with mixed water where we only have measurements of the final combined volume and the final stabilized temperature. Our goal is to solve the corresponding inverse problem: inferring the initial volume and temperature of both the hot water and the cold water sources that were used to fill the tub.
Notebook Cell
import cuqi
import numpy as np2.1. The forward model¶
Let be the volume of hot water, be the temperature of hot water, be the volume of cold water, and be the temperature of cold water. The forward map determines the final volume and the final temperature of the mixed water:
The final volume is simply the sum of the hot and cold water volumes:
The final temperature is calculated as the weighted average of the two temperatures, based on their respective volumes:
def forward_map(h_v, h_t, c_v, c_t):
# volume
volume = h_v + c_v
# temperature
temp = (h_v * h_t + c_v * c_t) / (h_v + c_v)
return np.array([volume, temp]).reshape(2,)Advanced MCMC algorithms, such as the MALA or HMC, rely on gradient information to efficiently explore the target distribution. Specifically, they need the gradient of the log-posterior, which in turn requires the derivative of the forward model with respect to each input parameter.
By supplying explicit analytical gradients for , , , and , we make our sampler both significantly faster and numerically more stable compared to relying on finite-difference approximations.
Below, we define the action of the Jacobians (the directional derivatives) for each input parameter:
# Define the gradient with respect to h_v
def gradient_h_v(direction, h_v, h_t, c_v, c_t):
return (
direction[0]
+ (h_t / (h_v + c_v) - (h_v * h_t + c_v * c_t) / (h_v + c_v) ** 2)
* direction[1]
)
# Define the gradient with respect to h_t
def gradient_h_t(direction, h_v, h_t, c_v, c_t):
return (h_v / (h_v + c_v)) * direction[1]
# Define the gradient with respect to c_v
def gradient_c_v(direction, h_v, h_t, c_v, c_t):
return (
direction[0]
+ (c_t / (h_v + c_v) - (h_v * h_t + c_v * c_t) / (h_v + c_v) ** 2)
* direction[1]
)
# Define the gradient with respect to c_t
def gradient_c_t(direction, h_v, h_t, c_v, c_t):
return (c_v / (h_v + c_v)) * direction[1]In CUQIpy, geometries govern how parameters are interpreted and presented. Because we have four distinct inputs with completely different meanings (two are volumes, two are temperatures), it is natural to represent each of them as a Discrete geometry. Then we define a tuple for our domain_geometry consisting of the 4 Discrete geometries. Similarly, the range_geometry defines the physical structure of our output space: exactly two discrete point measurements (temperature and volume).
domain_geometry = (
cuqi.geometry.Discrete(['h_v']),
cuqi.geometry.Discrete(['h_t']),
cuqi.geometry.Discrete(['c_v']),
cuqi.geometry.Discrete(['c_t'])
)
range_geometry = cuqi.geometry.Discrete(['temperature','volume'])Now we wrap everything together into a cuqi.model.Model object. The constructor simply requires:
forward: our Python function simulating the mixtureforward_map.gradient: the tuple of our four analytically computed gradients.domain_geometry: our tuple defining the four input parameters.range_geometry: our geometry specifying the two outputs.
When initialized, this object internally coordinates evaluating subsets of inputs and parsing gradients on demand during the MCMC sampling process.
model = cuqi.model.Model(
forward=forward_map,
gradient=(gradient_h_v, gradient_h_t, gradient_c_v, gradient_c_t),
domain_geometry=domain_geometry,
range_geometry=range_geometry
)2.2. Prior distributions¶
In real-world applications, prior distributions should be carefully elicited based on physical constraints and expert knowledge. However, for the purpose of this pedagogical example, we choose simple, somewhat arbitrary priors to illustrate the workflow in CUQIpy. Specifically, for the fluid volumes and the hot water temperature, we utilize Uniform distributions to reflect simple bounds. For the cold water temperature, we apply a TruncatedNormal distribution to demonstrate how more informative priors with specific boundaries can be incorporated.
h_v_dist = cuqi.distribution.Uniform(0, 60, geometry=domain_geometry[0])
h_t_dist = cuqi.distribution.Uniform(40, 70, geometry=domain_geometry[1])
c_v_dist = cuqi.distribution.Uniform(0, 60, geometry=domain_geometry[2])
c_t_dist = cuqi.distribution.TruncatedNormal(
10, 2**2, 7, 15, geometry=domain_geometry[3]
)2.3. Data, joint and posterior distribution¶
Then we construct the following objects:
Data Distribution: We assume the measurements of volume and temperature are corrupted by independent Gaussian noise. The
Modelwe defined serves as the mean of this distribution.Joint Distribution: We combine the data distribution and the priors into a single joint distribution.
Posterior: By conditioning the joint distribution on our observed data, we obtain the posterior distribution of our unknown parameters.
data_dist = cuqi.distribution.Gaussian(
mean=model(h_v_dist, h_t_dist, c_v_dist, c_t_dist),
cov=np.array([[1**2, 0], [0, 0.5**2]])
)
joint_dist = cuqi.distribution.JointDistribution(
data_dist,
h_v_dist,
h_t_dist,
c_v_dist,
c_t_dist
)
# Assume measured volume is 60 gallons and measured temperature is 38 degrees
# celsius
posterior = joint_dist(data_dist=np.array([60, 38]))2.4. Sampling with HybridGibbs¶
We will use CUQIpy’s HybridGibbs sampler. Gibbs sampling involves iteratively drawing samples from the conditional distribution of each parameter given the current state of the others. To do this in CUQIpy, we must define a sampling strategy: a dictionary that maps each of our target distributions to a specific MCMC sampler.
Here, we assign the MH sampler for the first 3 fields and the gradient-aware MALA sampler for the last field. As with the priors, this configuration is chosen primarily to demonstrate the flexibility of CUQIpy’s HybridGibbs framework which allows mixed sampling strategies. In a demanding real-world scenario, the choice of samplers would be carefully matched to the dimensionality and gradient availability for each parameter.
sampling_strategy = {
"h_v_dist": cuqi.sampler.MH(
scale=10.0, initial_point=np.array([30])),
"h_t_dist": cuqi.sampler.MH(
scale=10.0, initial_point=np.array([50])),
"c_v_dist": cuqi.sampler.MH(
scale=10.0, initial_point=np.array([30])),
"c_t_dist": cuqi.sampler.MALA(
scale=0.6, initial_point=np.array([10])),
}Then create the sampler and sample the posterior distribution.
np.random.seed(0)
cuqi.config.PROGRESS_BAR_DYNAMIC_UPDATE = False
hybridGibbs = cuqi.sampler.HybridGibbs(
posterior,
sampling_strategy=sampling_strategy)
hybridGibbs.warmup(1000)
hybridGibbs.sample(5000)
samples = hybridGibbs.get_samples()Warmup: 0%| | 0/1000 [00:00<?, ?it/s]Warmup: 100%|██████████| 1000/1000 [00:09<00:00, 100.83it/s]
Sample: 0%| | 0/5000 [00:00<?, ?it/s]Sample: 100%|██████████| 5000/5000 [00:52<00:00, 94.61it/s]
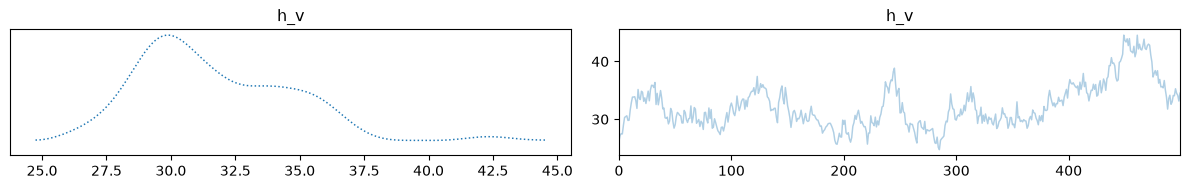
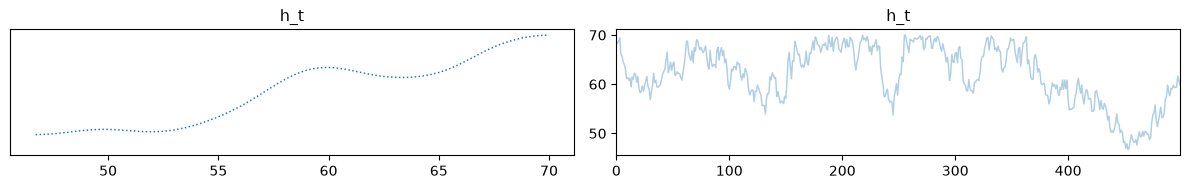
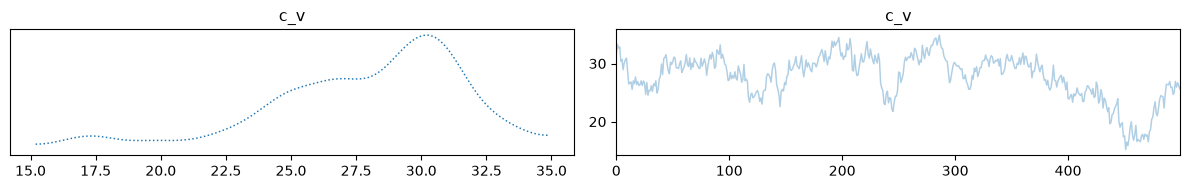
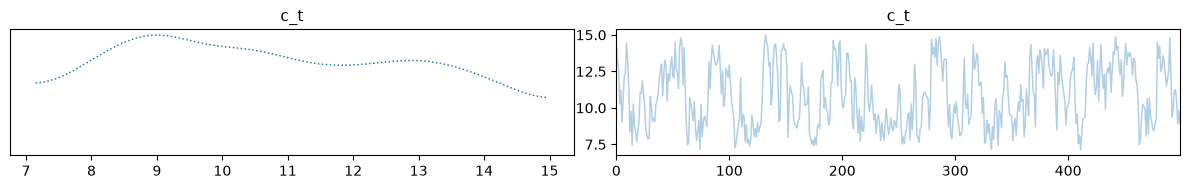
Finally, we can visualize our samples. Because we are trying to infer 4 initial state parameters from only 2 macroscopic measurements, the inverse problem is highly underdetermined. As we can see, the temperature and volume of the mix computed using the posterior means match the true solution very well.
# Compute mean values
mean_h_v = samples['h_v_dist'].mean()
mean_h_t = samples['h_t_dist'].mean()
mean_c_v = samples['c_v_dist'].mean()
mean_c_t = samples['c_t_dist'].mean()
# Print mean values
print(f"Mean h_v: {mean_h_v}, Mean h_t: {mean_h_t}, Mean c_v: {mean_c_v}, Mean c_t: {mean_c_t}")
print("Measured volume:", 60)
print("Mean predicted volume:", mean_h_v + mean_c_v)
print()
print("Measured temperature:", 38)
print("Mean predicted temperature:", (mean_h_v * mean_h_t + mean_c_v * mean_c_t) / (mean_h_v + mean_c_v))
# Plot trace of samples
samples['h_v_dist'].burnthin(1000,10).plot_trace();
samples['h_t_dist'].burnthin(1000,10).plot_trace();
samples['c_v_dist'].burnthin(1000,10).plot_trace();
samples['c_t_dist'].burnthin(1000,10).plot_trace();Mean h_v: [32.19099917], Mean h_t: [62.02483271], Mean c_v: [27.80258829], Mean c_t: [10.79161036]
Measured volume: 60
Mean predicted volume: [59.99358746]
Measured temperature: 38
Mean predicted temperature: [38.28202538]