The CUQIpy User Showcase repository is a dedicated space for collecting and sharing a diverse array of practical applications built using CUQIpy. The repository originated as a community-driven initiative where enthusiastic CUQIpy users generously contribute their own use cases, custom implementations, and experiments. Its main purpose is to serve as an educational and practical resource, providing real-world examples of how CUQIpy can be used to solve complex inverse problems and quantify uncertainty across different domains.
The repository houses a wide variety of solved problems, demonstrating the flexibility of CUQIpy. Examples include 1D and 2D deblurring/deconvolution (utilizing Besov priors, implicit priors, and smoothed Laplace methods), fusion plasma tomography, super-resolution imaging, and physical system modeling like spring systems and inverse Robin boundary problems. Users have also contributed examples that showcase advanced techniques such as plug-and-play priors (combining neural network denoisers with Bayesian inference), delayed acceptance with ODEs, dimension-reduction with Laplace priors, and creating custom forward models and samplers.
For example in the super-resolution imaging case study contributed by Rafael Flock, a novel method is proposed to perform uncertainty quantification for stochastic optical reconstruction microscopy (STORM). STORM, shown in Figure 1, is a super-resolution microscopy method based on single-molecule stochastic switching, where the goal is to detect molecule positions in live cell imaging. The noisy image is obtained by a microscope detecting the photon count of the (fluorescence) photoactivated molecules.
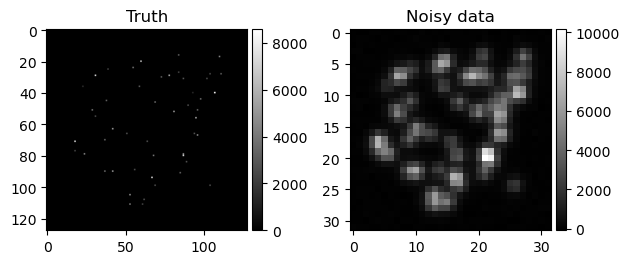
Figure 1. Ground truth and noisy data in a STORM problem. The goal is infer the location of molecules (white points in the left plot) from the noisy data shown in the right plot.
For the STORM problem, Laplace priors are an appealing choice for their ability to promote sparsity and handle heavy-tailed distributions, but their application in high-dimensional inverse problems is challenging. This case study proposes a novel approach that combines the method of Certified Coordinate Selection (CCS) with a gradient-based sampling method. CCS allows one to efficiently identify the components in the unknown parameters that contribute most to the update from the prior to the posterior, effectively reducing the dimensionality of the problem considered during inference, as shown in Figure 2.
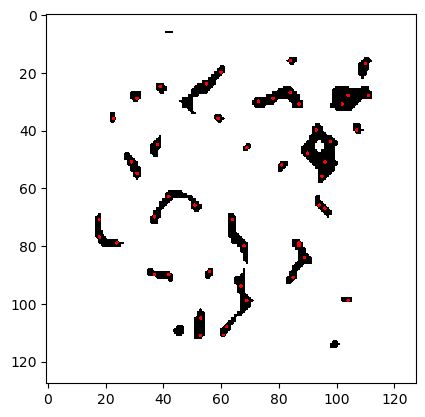
Figure 2. The selected coordinates after CCS are illustrated as black pixels and the exact locations of the molecules as red.
The CUQIpy developer team highly encourages new submissions and a detailed instruction on how to contribute is available at the User Showcase GitHub repository.
Resources¶
User-Showcase GitHub repository: https://
github .com /CUQI -DTU /CUQIpy -User -Showcase